Stata Tips #10 - Bayesian analyses in Stata 15
In this post, we are going to introduce the bayes and bayesmh commands. bayes is a prefix that you can attach to any of 45 existing Stata estimation commands to fit those models in a Bayesian framework. For example, if you have a logistic regression you run with logistic, you can now type bayes: logistic… and get the same model with prior information and Bayesian interpretation.
We’ll assume that you know what Bayesian analysis is and why you might want to use it, but if you would like to learn more, we recommend these resources (in order of increasing depth and complexity):
- Rasmus Bååth’s three videos
- StataCorp’s introductory videos on Bayes
- John Thompson’s book "Bayesian Analysis with Stata” (note that this pre-dates Stata 14 and so does not include bayes and bayesmh)
- As a comprehensive reference and a book to learn from, “Bayesian Data Analysis” by Andrew Gelman and friends.
Let’s start with bayes. In general, bayesmh is more flexible but also requires you to write out a lot more detail for your model. If you are fitting something fairly standard, you can save yourself the effort (and the scope for error) using bayes instead.
A simple example with bayes
The dataset impact.dta (downloadable here) contains details of 518 participants in a clinical trial of drug addiction treatments. They were randomly allocated to different durations of residential treatment at two sites and followed up for 610 days, so this will be a survival analysis. We’ll look at the effect of duration on relapse or loss to follow up, which are combined in this dataset. We’ll also adjust for site, but apart from that it is an over-simplistic analysis, used here just to illustrate the use of Stata. Don’t take it as medical evidence or base any decisions about addiction treatment on it!
We’ll use a Weibull proportional hazards survival model with the Stata command streg. If you are not familiar with these, don’t worry. The only thing you need to know is that it will produce regression coefficients, one for site B compared to site A, and one for long duration compared to short duration. Those coefficients will be log-hazard ratios, which are like risk ratios. A negative coefficient for duration would indicate that long treatment helps by reducing the hazard of relapse / loss to follow up.
We’ll look at the usual question of whether the confidence interval for long duration crosses zero or not, as this tells us whether we can call any benefit “significant”. But we’ll also ask a Bayesian question. The healthcare organization says that they can only afford long treatment if it reduces relapse / loss to follow up by more than 15%. What is the probability that the hazard ratio is below 0.85?
This sort of probabilistic statement, forbidden in frequentist statistics, is often what is needed for managerial and financial decisions.
Let’s get the data, tidy up the time variable, and define the survival with stset:
use "impact.dta", clear
replace time=610 if time==.
stset time, failure(relapse)
Fit the standard Weibull model:
streg i.treat i.site, distribution(weibull) nohr
And now the same thing with Bayes:
bayes: streg i.treat i.site, distribution(weibull) nohr
That’s exactly the same line, just adding bayes: to the beginning! The difference here is that there are prior distributions for the parameters and it is estimated using simulation. We left the priors as defaults, and you can see what those are above the results table in the output:
Model summary
------------------------------------------------------------------------------
Likelihood:
_t ~ streg_weibull(xb__t,{ln_p})
Priors:
{_t:1.treat 1.site _cons} ~ normal(0,10000) (1)
{ln_p} ~ normal(0,10000)
------------------------------------------------------------------------------
(1) Parameters are elements of the linear form xb__t.
The log-hazard ratios for treat (duration) and site both get normal distribution priors with mean 0 and variance 10,000 (standard deviation 100). That’s what we might call “diffuse” or “vague” and around any plausible value, very close to being flat.
You’ll notice that the Bayesian version takes notably longer, and that’s because it works by simulating parameter values many times (by default, 12,500, with the first 2500 getting discarded as “burn-in”). The coefficient estimates from the two methods are similar. In both cases, long duration looks beneficial, and the 95% confidence interval or credible interval is all on the good side of zero, but it does get close to zero.
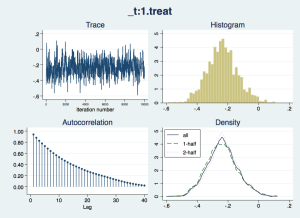
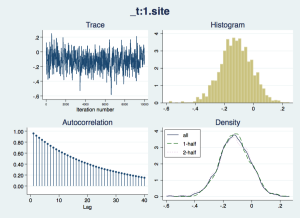
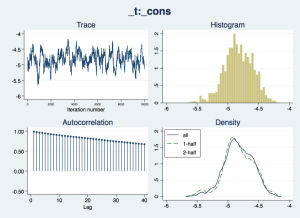
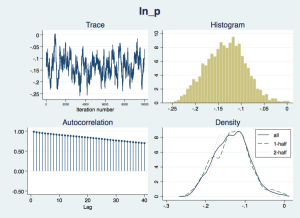
Now we should check that the “chain” of simulated draws from the posterior distributions – which inform the estimates and inferences in the results – look wel-behaved:
bayesgraph diagnostics _all
The traces appear to consistently explore the same range of values – they are not drifting slowly up or down – but there is considerable autocorrelation, suggesting that they are slow to move around the posterior distribution, especially for the nuisance parameters cons and ln_p. We would like more posterior draws to smooth out any bumps in the distribution that arise from this slow exploration, and we can do this with the mcmcsize option. We'll also get Stata to display dots to show us how it is progressing, with the dots option.
bayes, mcmcsize(50000) dots(1000): streg i.treat i.site, distribution(weibull)
This looks better; the “mixing” of the chain is just a little slow. There are options for thinning but we will side with Christian Robert (Chapter 2, “Handbook of MCMC”, CRC Press) and keep them all. Another way of assessing how slow mixing reduces the information available in the simulation is by using the effective sample size: how many independent samples would give you as much insight into the posterior distribution as you have from the current chain.
bayesstats ess
Efficiency summaries MCMC sample size = 50,000
bayesstats ess
Efficiency summaries MCMC sample size = 50,000
----------------------------------------------------
| ESS Corr. time Efficiency
-------------+--------------------------------------
_t |
treat |
long | 2696.99 18.54 0.0539
|
site |
B | 1172.09 42.66 0.0234
_cons | 230.69 216.74 0.0046
-------------+--------------------------------------
ln_p | 216.88 230.55 0.0043
----------------------------------------------------
We may be generating 50,000 draws from the posterior, but the autocorrelation means this is equivalent to 2697 independent samples in the duration coefficient, and as few as 217 in the nuisance parameters. We might even want more than 50,000 draws, but we’ll leave it for now. It would also be a good idea in a serious analysis to try different initial values and priors, just in case there is some sensitivity to either of these.
Now we can find the probability of the hazard ratio for long treatment being under 0.85 (i.e. {_t:1.treat} < -0.1625).
bayestest interval {_t:1.treat}, upper(-0.1625)
Interval tests MCMC sample size = 50,000
prob1 : {_t:1.treat} < -0.1625 ----------------------------------------------- | Mean Std. Dev. MCSE -------------+--------------------------------- prob1 | .77768 0.41581 .0081723 -----------------------------------------------
That’s a 78% probability of the longer treatment reaching the affordability threshold the organization is looking for.
Another way we can investigate the joint posterior distribution of the parameters is by saving the draws into a new data file, using the saving option.
bayes, mcmcsize(50000) dots(1000) saving(impact_bayes, replace): ///
streg i.treat i.site, distribution(weibull)
Any analyses you then want to carry out need to be weighted by the _frequency variable. This is because Bayesian simulation is based on Markov chains, which sometimes move to new values, and sometimes stay where they are, repeating the same values.
use impact_bayes.dta, clear
summarize [fw=_frequency]
Or you can expand the data to its full size (50,000):
expand _frequency
sort _index
generate newindex = _n
You might wonder what the eq* variable names mean, and they appear in the order that parameters are listed in the macro e(parnames):
display e(parnames)
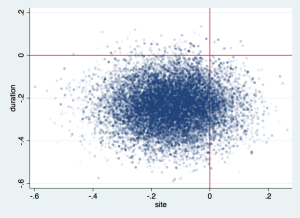
Now you can calculate further statistics as combinations of parameters at each draw, and then summarise those, or just look at the joint probabilities with a scatterplot:
scatter eq1_p2 eq1_p4, msize(vsmall) mlalign(outside) mlwidth(none) mcolor(%6) ///
xtitle(site) ytitle(duration) xline(0) yline(0)
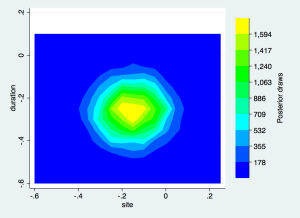
or a contour plot:
gen sitecat=floor(eq1_p4*20)/20
gen duracat=floor(eq1_p2*20)/20
collapse (count) n=newindex, by(sitecat duracat)
twoway contour n duracat sitecat, levels(10) scolor(blue) ecolor(yellow) ///
xtitle(site) ytitle(duration) ztitle("Posterior draws")
Specifying priors
So far, we’ve just used default priors. We might have some external information from another study on the same population, or we might just want to prevent the simulation from straying into values we feel are impossible and will cause computational problems. Specifying bespoke priors is simple, and essential for good analyses. Let’s take a simple example with the auto dataset.
sysuse auto, clear
regress price mpg
bayes: regress price mpg
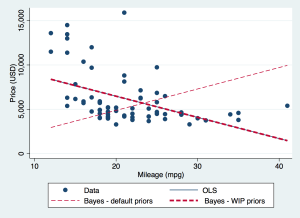
Ordinary least squares regression tells us that the coefficient of MPG predicting price is -238.9, with standard error 53.1. The corresponding Bayesian analysis with default priors gives 240.4 with standard deviation of the posterior draws 20.1. This has flipped from a large negative to a large positive value!
What’s going on? Let’s look at the whole regression equation. From OLS:
pricei = 11253.1 - 238.9mpgi + ei
where e ~ N(0, 2623.7)
Those are some big numbers. The price is in US dollars, of course.
and from default prior Bayes:
pricei = 47.9 + 240.4mpgi + ei
where e ~ N(0, 4001.7)
Recall that the default prior for the regression coefficient is normal with mean 0 and standard deviation 100. That prior is going to return an extremely low probability for the intercept coefficient of 11253.1. In fact, because of digital rounding, it will return a probability of zero (try display normalden(11253.1/100)), and prevent the simulation from going there at all, and with the intercept under-estimated, the slope of MPG has to be over-estimated to hit the data at all.
So, we have learnt the hard way that there is no such thing as a non-informative prior. What is non-informative in one context is highly informative in another. Andrew Gelman said, “sometimes classical statistics gives up. Bayes never gives up, so we’re under more responsibility to check our models.”
Instead of the defaults, let’s think through what to use as a weakly informative prior (WIP). These aim to keep the computation to a plausible range but impose no more than that.
We know that the MPGs are in the tens, and the price in the tens of thousands, so we expect a slope of perhaps a thousand or more. Let’s use a normal distribution with mean 0 and standard deviation of 2000 – that’s a variance of 4,000,000. Then, we could also use uniform distributions for the intercept, covering the range of prices with an extra 10,000 either side (because the y-axis of zero miles per gallon is not in the data so there is some extrapolation). For the residual variance, let’s allow it to be as high as 5000, and use a uniform distribution, but we’ll exclude very small variances with a lower limit of 0.1. This just avoids computational problems that sometimes arise when Bayesian simulations get into very small variances.
All of these can be included quite intuitively with repeated uses of the prior option. We’ll also set sensible initial values with the initial option.
bayes, prior({price: mpg}, normal(0, 4000000)) ///
prior({price: _cons}, uniform(-10000,25000)) ///
prior({sigma2}, uniform(0.1, 5000)) ///
initial({price: _cons} 12000 {price: mpg} -300 {sigma2} 500) ///
: regress price mpg
This gives a solution very close to the OLS regression, as seen in the graph.